-
Address:
17888 67th Court North
Loxahatchee, FL
-
Mail us:
contact@wrightacademia.org
- submit manuscript
Table of content
Review Article |
Open Access |
Volume 2 | Issue 1 |
Hurdles and Directions in the Fight against COVID-19
Sulagna Mandal1, Debprasad Chattopadhyay2,3, Ishwar Singh2 and Aparna Mukhopadhyay1
1Department of Life Sciences, Presidency University, Kolkata, India
2ICMR-National Institute of Traditional Medicine, Nehru Nagar, Belagavi, India
3ICMR-National Institute of Cholera and Enteric Diseases, Kolkata, India
*Corresponding author: Aparna Mukhopadhyay, Department of Life Sciences, Presidency University, 86/1 College Street, Kolkata-700073, India, Email: aparna.dbs@presiuniv.ac.in
Citation: Mandal S, Chattopadhyay D, Singh I, Mukhopadhyay A (2021) Hurdles and Directions in the Fight against COVID-19. J SARS-CoV-2 COVID 2:004.
Copyright © Mandal S, et al.
Received: |
Accepted: |
Published: |
COVID-19 has ushered in a new decade with potential to leave a permanent mark in the history of mankind. While some vaccine candidates are in circulation, we are still largely ignorant on SARS-CoV-2. This review summarizes information on the SARS-CoV-2 as revealed by the innumerable publications that have emerged. Parallels have been drawn with the more studied predecessors SARS-CoV and MERS, while the areas that still pose challenges in vaccine, prophylactic or therapeutic development have been highlighted. Thus, this review attempts to provide a holistic understand of the virus along with possible areas where successful therapeutic development can proceed.
Introduction
With an alarming and rising number of morbidity and mortality, the COVID-19 pandemic has made a permanent mark on the history of mankind/human evolution. The disease originally emerged from Wuhan, China at the end of December, 2019, has wiped out over 2.7 million of world population and infected over 124 million globally, as of March 23rd, 2021, as per WHO reports. Owing to frequent international travel this 160 nm pathogen could not be contained; rather transmitted from country to country, rapidly infecting more than 200 countries across the seven continents, as per WHO. Living in an era of advanced science and technology, it was unimaginable that a tiny ultramicroscopic (160 nm) nucleoprotein particle could wreck an unprecedented havoc to bring the entire world to a standstill, crashing the global economy, affecting the livelihood and careers of millions.
Although, coronaviruses has been studied since mid-1960 [1], it gained prominence during the Severe Acute Respiratory Syndrome (SARS) outbreak in China (November 2002-July 2003) [2] followed by Middle-East Respiratory Syndrome(MERS) outbreak (2012-2015) in the Middle-East countries [3]. While researchers were still pursuing studies on SARS, underlying mechanisms of pathogenicity of MERS and investigating a tool that would either act as a pan-coronavirus inhibitor or a novel therapy that would efficiently target SARS- and MERS-CoV, the highly pathogenic variant (initially named as 2019-nCoV by WHO) emerged. Among the seven human coronaviruses, namely, 229E, NL63, OC43, HKU1, SARS, MERS and SARS-CoV-2, the newest and the seventh member, SARS-CoV-2 has the most sequential resemblance to bat-derived SARS-like coronaviruses with 88-89% similarity followed by SARS (79.5%) [4]. Like its predecessors SARS-CoV and MERS-CoV, SARS-CoV-2 have shown different degrees of pathogenicity and virulence, such as, bronchitis, pneumonia with renal involvement [5], however proving to be most virulent among the three. COVID-19 is associated with a broad range of clinical manifestations ranging from asymptomatic pneumonia to severe acute respiratory diseases, enteric, hepatic and neurologic diseases [6]. Patients with co-morbidities and immuno-incompetence have the worst prognosis [7]. While SARS infects ciliated bronchial epithelial cells and type-II pneumocytes through angiotensin-converting enzyme 2 (ACE2), MERS infects un-ciliated bronchial epithelium and type-II pneumocytes through dipeptidyl peptidase 4 (DPP4) receptor, also known as CD26 [8,9]. SARS-CoV-2, just like its predecessor SARS-CoV also uses ACE2 as cellular receptor [10].
The urgency to develop specific drugs and vaccines which started since SARS outbreak, gained prominence and boost since the COVID-19 pandemic started. Accelerated efforts were aimed towards therapies which would either show potent antiviral activity against pan-coronaviruses or be targeted against SARS-CoV-2, thereby lowering the severity of the pandemic and also trying to eliminate the probability of another outbreak. Although, wait for vaccine seems to be over, it is too soon to arrive at the conclusion about potency and safety of these vaccine candidates. In this review, we focus on the molecular virology of SARS-CoV-2 along with possible hindrances faced by scientists in the venture of drug development to fight the pandemic with respect to its transmission and mutation of the virus. Further attention is drawn towards viral immunology and mechanism of pathogenesis that evolved to evade the most crucial antiviral immune response. Therefore, this review might provide new direction in therapeutic development against COVID-19 by furthering our knowledge on the above mentioned aspects of SARS-CoV-2.
Molecular Virology
Structure and Genome Organization of SARS-CoV-2
Belonging to the order Nidovirale, family Coronaviridae, SARS-CoV-2 is an enveloped virus containing non-segmented positive sense, single strand RNA genome of ~30 kilobases (Figure 1) [11-13]. Coronaviruses are divided into 4 genera, α-, β-, γ-, and δ-coronaviruses. The β-Coronaviruses are further divided into A, B, C and D lineages. Among the seven identified human coronaviruses so far, SARS-CoV and SARS-CoV-2 belong to the β-coronaviruses lineage B [14].
The term "corona" was derived from the appearance of a crown-like spike projecting from the surface of the envelope. All coronaviruses share the common features in the organization and expression of their genome, a very large open reading frame 1 (ORF 1), comprising about the two-third of the genome that encodes two large overlapping ORF1a and ORF1b as depicted in (Figure 1). Other ORFs encode the structural proteins. Additionally, an ORF8 is found to exist in SARS-CoV and SARS-CoV-2, whose exact function is unknown, but is predicted to facilitate host shifts [15].
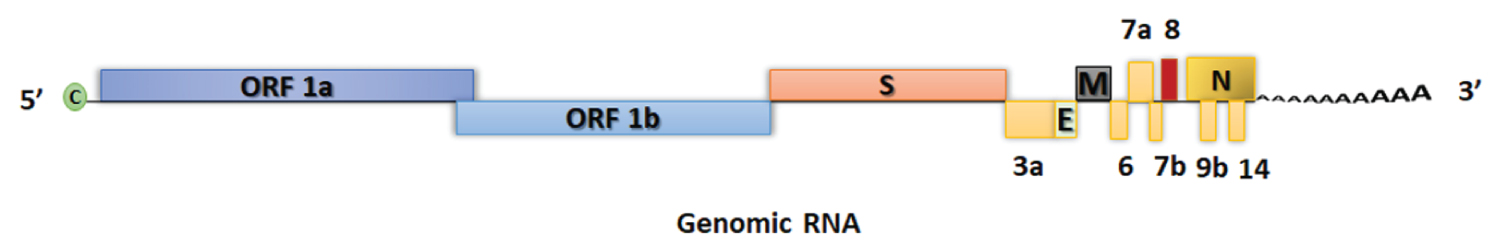
Figure 1: Genomic Organisation of SARS-CoV-2.
The ~30 kb RNA genome encodes for two overlapping ORFs (1a and 1b). These are translated to produce two large polyproteins pp1a and pp1b. The rest of the genome is transcribed into a nested set of subgenomic mRNAs. These viral polyproteins are further processed by virally encoded cysteine proteases, papain-like protease (PLpro) and 3-chymotrypsin-like protease (3CLpro). Sixteen (16) non-structural proteins (nsp1-nsp16) are encoded by ORF 1a/b at the 5' end, followed by structural proteins spike (S), membrane (M), envelope (E) and nucleocapsid (N), all of which are encoded by ORFs at the 3' end. Each structural protein is depicted by its single letter code. Numbers indicate the non-structural elements.
Both structural and non-structural proteins play a very crucial role in the viral life cycle as well as pathogenesis, depicted pictorially in (Figure 2) [16-18]. Therefore, these proteins always serve as potent targets for antiviral therapy. Primary candidates include the Spike (S) protein which forms the surface 'corona' of the coronavirus, as it's primarily responsible for entry and eliciting an adaptive antiviral response. Other targets include RdRp and nsp3 which are essential proteins involved in the replicase complex. Presence of viroporins encoded by ORF3a is unique to SARS-CoV and SARS-CoV-2 suggested by structural analysis [19,20]. These accessory proteins form membrane pores which regulate cellular ion-channels, which make them invaluable tools for anti-viral therapy and need to be explored further.
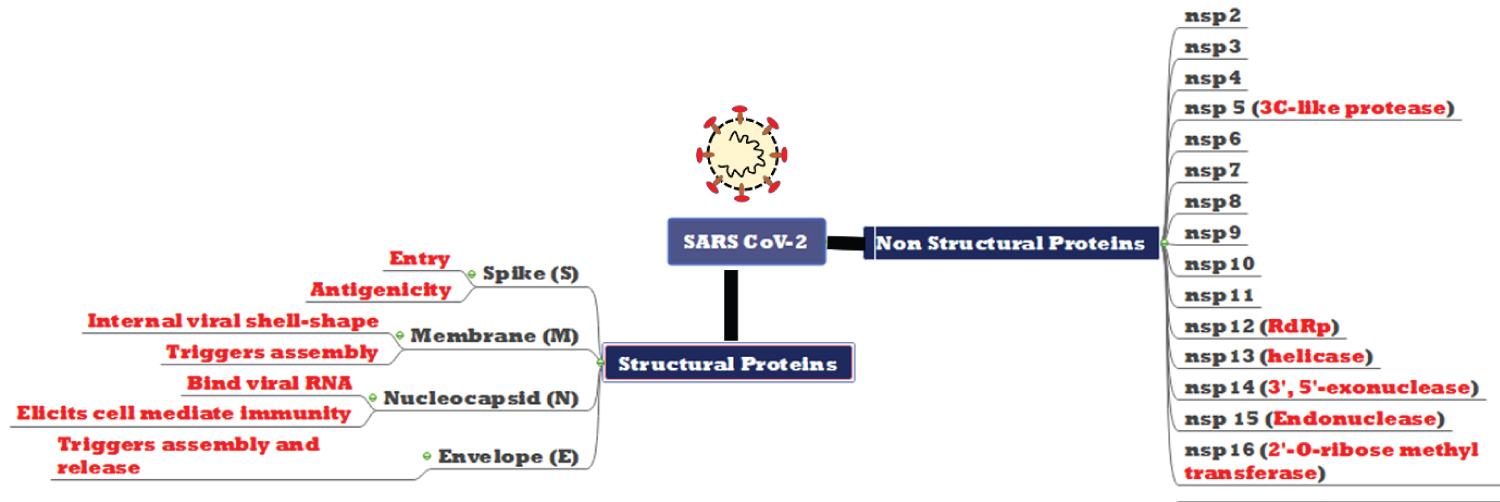
Figure 2: Structural and non-structural proteins encoded by SARS-CoV-2.
The primary function of each protein is depicted in red. Function of all 16 non-structural proteins is not clear. ORF1b encodes the RNA dependent RNA polymerase (RdRp), helicase and an mRNA cap-1 methyl-transferase as predicted by computational analysis. An exo-ribonuclease is also encoded by nsp14 of ORF1b, which is a unique feature of coronaviruses, contributing to high replication fidelity. Among four structural proteins, the S protein is required for species specificity, receptor binding and fusion. The M protein is a central organizer of viral assembly and defines the shape of viral envelope together with E protein. However, the majority of the E protein is localized at the site of intracellular trafficking, where it is found to participate in viral assembly and budding. Apart from these proteins, the N protein is actively involved in complete virus formation.
Here we highlight features of SARS-CoV-2 in comparison to the relatively better studied SARS-CoV (Table 1). Innumerable publications so far have focused on the Spike protein (because of its obvious importance in host entry and innate immune response) and many of the structural peculiarities have been well documented. Structural analysis reveals that the spike (S) protein of SARS-CoV-2 is closely related to SARS-CoV, sharing about 89.8% sequence similarity in their S2 subunit [14], an important target of fusion inhibitors at present. This type-I glycoprotein binds to the cellular receptor Angiotensin-Converting Enzyme-2 (ACE2), present in almost all major organs like lungs, heart, kidney and intestine [21]. The main physiological role of ACE2 is in the maturation of angiotensin, a peptide hormone involved in regulation of vasoconstriction and blood pressure [22]. The S protein of both SARS-CoV and CoV-2, has two extracellular structural domains S1 and S2. Receptor binding is mediated by receptor binding domain (RBD) of S1 subunit, followed by fusion via S2 subunit. After receptor binding, the S protein undergoes conformational changes followed by cleavage at its S1/S2 junction by the host cell protease-transmembrane protease serine 2 (TMPRSS2) [23]. This priming leads to destabilization of the prefusion structure, cleaving S1 subunit and transition of S2 subunit to a stable post-fusion conformation [24,25]. After binding through RBD, the heptad repeat 1 (HR1) and 2 (HR2) domains in S2 subunit interact and a six-helix bundle (6-HB) fusion core is formed, bringing viral and cellular membranes in close proximity to accelerate fusion and infection, documented by X-ray crystallography. Studies also suggest cathepsin-L driven processing of spike protein in endosomes, indicating receptor-mediated endocytic entry of both SARS-CoV and CoV-2 (Figure 3). Importance of other cellular proteins like PIKFYVE (Phophatidylinositol 3-phosphate 5-kinase), TPC2 (Two pore channel subtype 2) in target cells that largely influence the viral entry have been highlighted in a recent study [26,27]. However more in-depth knowledge is needed on exact viral entry mechanisms to develop preventives and could be the Achilles' heel faced by modern science in developing antivirals with concerted efficacy (Box 1).
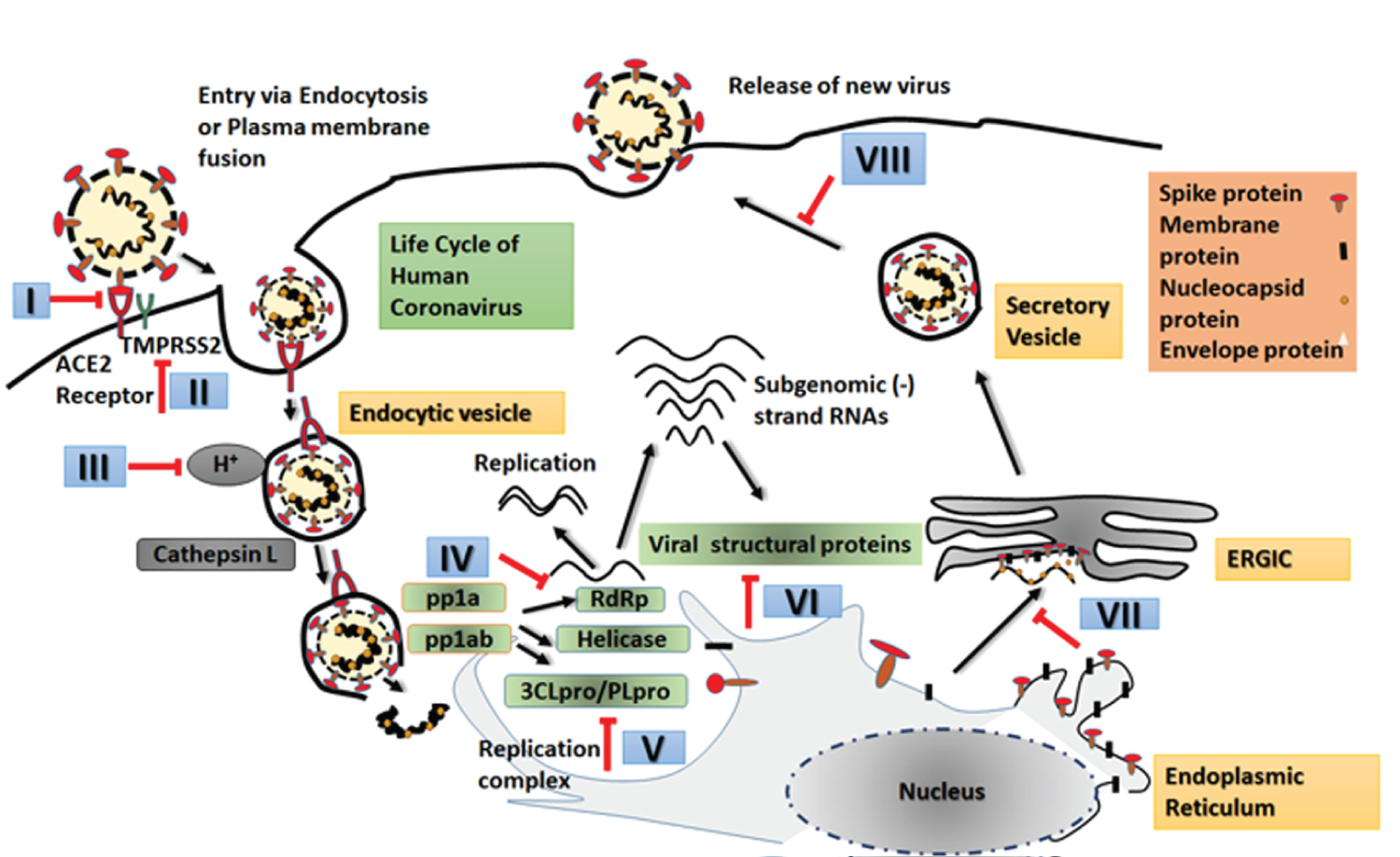
Figure 3: Life Cycle of SARS-CoV-2.
Life cycle begins immediately after binding of viral spike protein with host ACE2 receptor. Internalization by endosome. Release of viral genome into host cell cytoplasm followed by translation of ORF1a and ORF1b into two polyproteins (pp1a and pp1ab) by the host translational machinery. The pp1ab synthesis requires a -1 ribosomal frameshift but is formed from both ORFs. Further modifications occur when virally encoded proteases (3CLpro, PLpro) cleave these polyprotein to produce downstream proteins involved in viral replication in which a full length negative strand RNA is synthesized. Other downstream structural and non-structural proteins are synthesized via a nest of sub-genomic negative sense RNAs. Incorporation of structural proteins in endoplasmic reticulum (ER) followed by assembly in endoplasmic reticulum-Golgi intermediate compartment (ERGIC). Budding and release of new virus particles. Potential therapeutic targets are indicated by red arrows. Numbered steps are areas where intervention of the life cycle can be done for development of therapeutics. Some of them have been tapped and several are already in clinical trial as detailed in text.
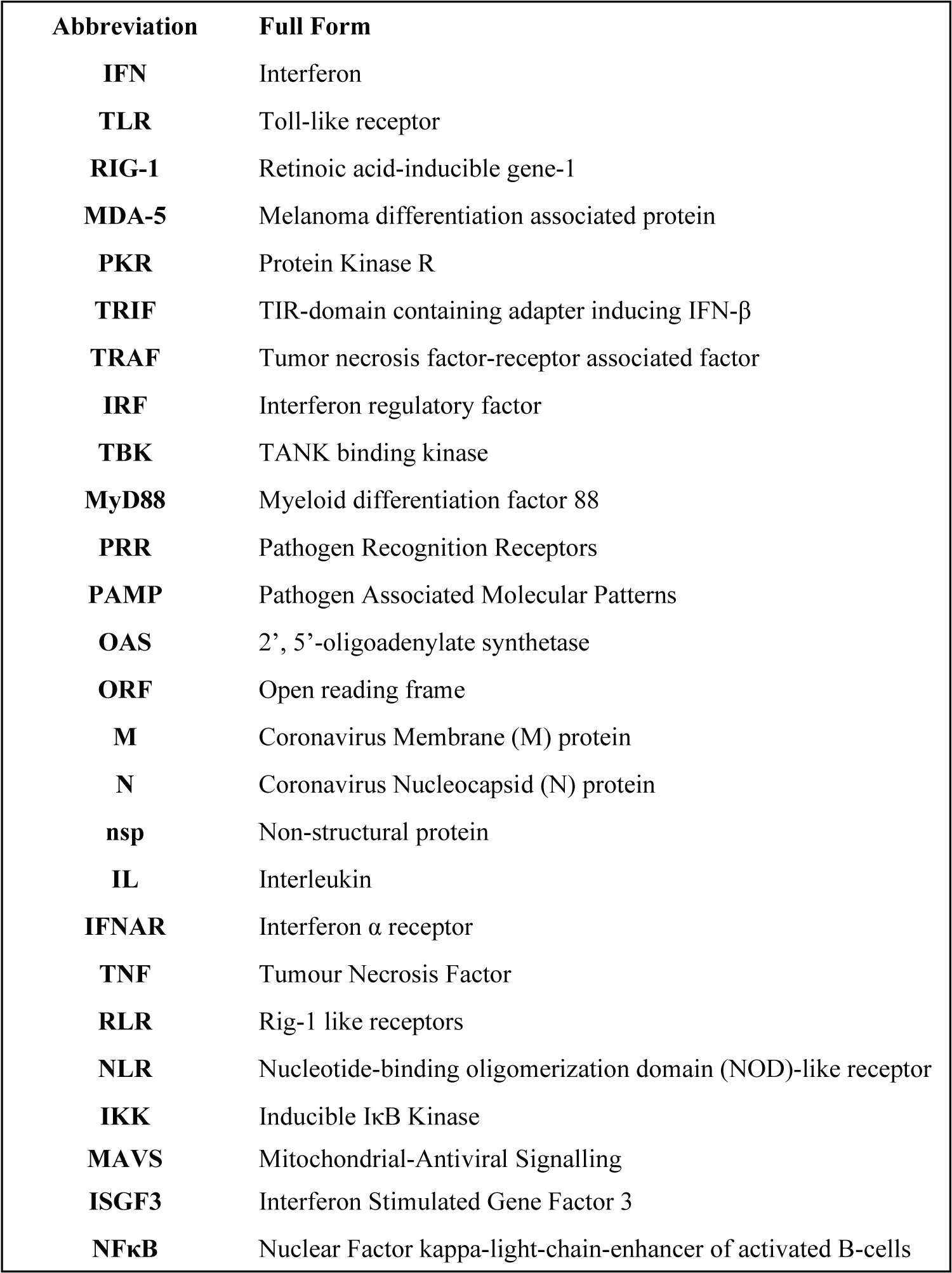

Box 1: List of Abbreviations and Full Forms.
Life cycle of human coronavirus: A challenge
An in-depth knowledge of virus biology is essential to therapeutic development (Figure 3) in this fight against COVID-19. The life cycle of the virus begins with viral entry (attachment fusion and penetration) into the target cell (goblet and ciliated cells in lungs and bronchioles) following membrane fusion, via Ca2+ dependent mechanism [32] or endocytosis, as described in (Figure 3). The complete virus is released from the host cell by fusion of virion-containing vesicles with the plasma membrane [33]. Virus replication possibly induces ER stress thus activating the unfolded protein response (UPR) signalling pathway [34]. Further investigations might make use of the UPR pathway as potent therapeutic target against emerging pathogenic human coronaviruses like SARS-CoV-2.
Transmission and Spread
Having originated from bats, SARS-CoV-2 has successfully crossed the species barrier and adapted human to human transmission, just like its predecessors [35]. The major route of transmission for SARS-CoV-2 include droplet transmission, aerosol of close room and perhaps feco-oral [36,37]. A possible nosocomial transmission is also involved, as evidence exists for past nosocomial infections of SARS [9]. The zoonotic source of this virus is still debatable, though phylogenetic analysis repeatedly relates it to bat-derived SARS-like coronaviruses (bat-SL-CoVZC45 and bat-SL-CoVZXC21) with 88-89% homology. The disease manifestation is reported in cats and tigers, but not in dogs and pigs, owing to low expression of ACE2 in the airways of dogs and pigs compared to the other susceptible species as well as homology of ACE2 between humans and cats [38]. Investigation of changes in ACE2 across species have identified many key changes between those that are infected vs. those that aren't. Consistent with studies related to SARS-CoV, the super spread of COVID-19 is also linked to surface stability and viability of SARS-CoV-2, mentioned in (Table 1). Higher binding affinity of SARS-CoV-2 S protein with host ACE2 indicates that the organs with high expression of ACE2 are the potential targets of SARS-CoV-2. Since most of the vital organs of the human body express ACE2, the morbidity and mortality rate is further enhanced as the virus can potentially infect multiple organs. Consistent with the wide spread of infection, we can assume the presence of more than one type of cellular receptor for viral entry besides ACE2. Hence, further studies should investigate the role of any other cellular receptor, if any, on viral pathogenesis. It is probable that this missing piece of knowledge might be a key to targeted therapeutics.
A major drawback in COVID research is the absence of an established animal model. A few recent studies claim that ferrets might be used as potential animal model to study viral pathogenesis, evidenced by very low titre of viral RNA in nasal turbinate, rectal swab and shedding of infectious virus in faeces. Another study also claims humanized mice to be a model to study viral pathogenesis. Infection in ferrets exhibit limitations due to the low viral titre in lungs than that exhibited by SARS-CoV and MERS-CoV infected hACE2 and hDPP4 transgenic mice, indicating that rigorous research is needed to establish an animal model [39,40]. Apart from these, Syrian hamsters may also serve as good model for SARS-CoV infection.
Mutation of SARS-CoV-2: Yet another obstacle
Mutations are constant, inevitable and an integral part of evolution for all life forms, especially in RNA viruses. Coronaviruses show varying mutation rates compared to other single-stranded RNA viruses. On an average coronaviruses have ~10-4 substitutions per year per site [5]. The hotspots of mutation involve regions such as ORF8, ORF3, spike protein and nsp3 [41]. Its large genome size contributes to extra plasticity in genome modifications by mutation and recombination which increases the probability of intra-species variability, interspecies host jump and emergence of novel coronaviruses under right conditions [42]. Although, occurrence of erroneous mutations during replication is comparatively lower in coronaviruses compared to other RNA viruses due to high proofreading activity carried out by 3'-5' exonuclease encoded by nsp14 [17], evolution of coronaviruses is in progress as evident by the newly emerging strains in UK, Brazil and other parts of the world. Increased recombination during RNA replication might also lead to mutations. Generation of nested set of sub-genomic mRNAs is also responsible for increased homologous recombination among closely related genes from different coronavirus lineages. Besides circulation in multiple hosts which serve as mixing vessels, is likely to increase the recombination events. Evidence suggests that SARS-CoV shares its recombination events with α and γ lineages. A number of specific breakpoints and small recombinant regions have been found in the RdRp (nsp12), parts of nsp9, nsp10 and nsp14, based on sequence analysis of different strains of SARS-CoV [43]. Studies have shown recombination between SARS-related bat-associated CoV generated an entirely new strain in civets with a breakpoint at the nsp16/spike and S2 region. Furthermore, SARS-CoV and SARS-CoV-2 [44] is shown to have acquired an ORF8 by gene gain mechanism that may facilitate host shift. Additionally, gene gain and gene loss leads to rapid evolution in these viruses [45]. Plasticity of SARS-CoV and other bat-derived SARS-like coronaviruses spike protein is yet another attributable factor to viral mutation. Multiple substitution events allow escape of the virus from antibody neutralization, while retaining receptor specificity. This flexibility allows usage of a wide range of zoonotic receptors to human ACE2 receptors [46].
Recent phylogenetic network of analysis suggests that ancestral SARS-CoV-2 has undergone mutations leading to different variants. Some studies classify them into various lineages namely A, B and C distinguished by amino acid changes [47]. Both synonymous and non-synonymous mutation has led to different lineages, as lineage B supposedly diverged from parent lineage A and C from B. Lineage A and C is speculated to be prevalent in the USA and European countries while B lineage is more prevalent in East-Asian countries. Another study classifies them into two major genotypes- type I and type II, which are further subdivided [48]. Type II emerged from type I and is prevalent. Genomic analysis reveals synonymous mutations in two sites, which is correlated to high translational efficiency in type II strains than type I, thus implying their rapid transmission and infection amplitude. Further genotyping showcases high frequency SNP mutation sites in the critical proteins like S protein, RNA polymerase, primase and nucleoprotein of SARS-CoV-2 [49]. Similar to its predecessors, several amino acid substitutions were identified in RBD of S protein of SARS-CoV-2, suggesting its changing tropism and increased pathogenesis [50]. Further evidence suggests involvement of many mutation events including deletion and mis-sense mutations in parts of the genome encoding the major structural and non-structural proteins, except envelope protein [51,52]. Two mutations reported in nsp6 might favour evasion of CoV-2 from cellular immunity, specifically autophagy, a critical stress-response pathway of the cell [53]. Autophagosomes are formed but lose their ability to deliver the viral components to lysosomes for degradation. Nevertheless new variants were also identified in India by in silico analysis [54]. Non-synonymous mutation was observed in nsp3 gene that might be related to its transmission rate. However, experimental validation is unavailable to support this in silico study. Evidences only suggest gradual evolution of SARS-CoV-2 leading to different variants of the original Wuhan strain. Among the new variants in circulation, D614G variant gained most prominence. Through its evolution in humans, the aspartate residue at 614 position got replaced by glycine at the carboxy terminal domain of S1 subunit of spike protein of SARS-CoV-2 and epidemiological study suggested higher transmission of the G614 variant [55]. Further investigation led researchers to claim higher replication of the new variant ex-vivo using primary human nasal epithelial (HNE) cells, large airway epithelial (LAE) cells and distal lung small airway epithelial (SAE) cells from multiple donors [56]. Besides, increased viral load and fitness of G614 has been suggested by few in vivo studies [57,58].
Attention should be drawn to the latest variant in circulation and mostly spoken of is the UK variant or B.1.1.7. Although, this variant is mostly predominant in the UK, it has been detected in several other countries including India, the USA and Japan [59]. Unlike the other variants, it has acquired multiple mutations in the spike protein including deletion 69-70, deletion 144, N501Y, A570D, D614G, P681H, T716I, S982A and D1118H [60]. Almost 70% higher transmission rate of the UK variant has been speculated, despite lack of experimental evidence. In light of exceptionally large number of mutations in the spike region, pathogenicity of this variant needs to be determined immediately.
High frequency of recombination as well as continuous interaction between multiple species of coronaviruses amongst animal host pool leads to emergence of pathogenic zoonotic coronaviruses and potential outbreaks already seen and may again happen in near future. The question that concerns scientists is whether these variants are more transmissible or more pathogenic. Emergence of these variants and their increasing prevalence have kept virologists busy in studying how these variants can lead to altered pathology, transmission, spread and prognosis of the disease. This also questions the efficiency of current vaccine candidates making the fight against this virus challenging.
Immune Response and Its Evasion
Commonly, patients exhibiting poor prognosis have been reported to experience a cytokine storm which brings about an overwhelming immune response that often results in a wide spread and uncontrolled inflammatory response in the body. In patients presenting acute respiratory syndrome (ARS) due to COVID-19, circulating levels of pro-inflammatory cytokines (IFNα, IFNγ, IL-1β, IL-6, IL-12, IL-18, Il-33, TNFα, TNFβ and chemokines CXCL10, CXCL8, CXCL9, CCL2, CCL3, CCL5 along with MCP-1 (monocyte chemoattractant protein 1) have been reported, similar to what has been reported in SARS and MERS infections [61-63]. Although many reports have extensively reviewed the pathology of a typical cytokine storm, especially in the context of SARS and MERS infections, details on COVID-19 are restricted to correlation between cytokine levels found in patients with mild symptoms to those requiring ventilation and scanty reports from autopsies [64,65]. This review briefly explains the basics of a cytokine storm in the context of COVID-19 as many extensive reviews are available. Cytokines are released from activated immune responsive cells which include interferons, interleukins, chemokines and TNF-α [66,67]. Of these interferons offer the first line of defence against viral infection and have been discussed below.
Interferon signaling system
As depicted in (Figure 4), abbreviations stated in Box 1 the innate immune response is triggered by the presence of viral RNA that acts as PAMPs to activate specific PRRs such as TLR-3 and TLR-7. Upon activation of the PRRs, numerous signal transduction pathways are activated, which together leads to activation of interferon genes. Type I and II interferons upon release bind to their receptors on specific cells to trigger the formation of ISGF3 that act as transcription factor to initiate synthesis of numerous products that would initiate an antiviral response by promoting immunomodulatory, antiviral and anti-proliferative activity. IFN signalling primarily acts through the JAK-STAT pathway. Amongst others, such products produced as a result of IFN signalling include synthesis of 2', 5'-oligoadenylate synthase (OAS) that cleaves viral ds and ssRNA via activation of the OAS/RNase L pathway. Other effects include activation of NFκB leading to promotion of T-cell differentiation, NKC activation, DC maturation, and apoptosis of the infected cell and even of the bystander uninfected cell.
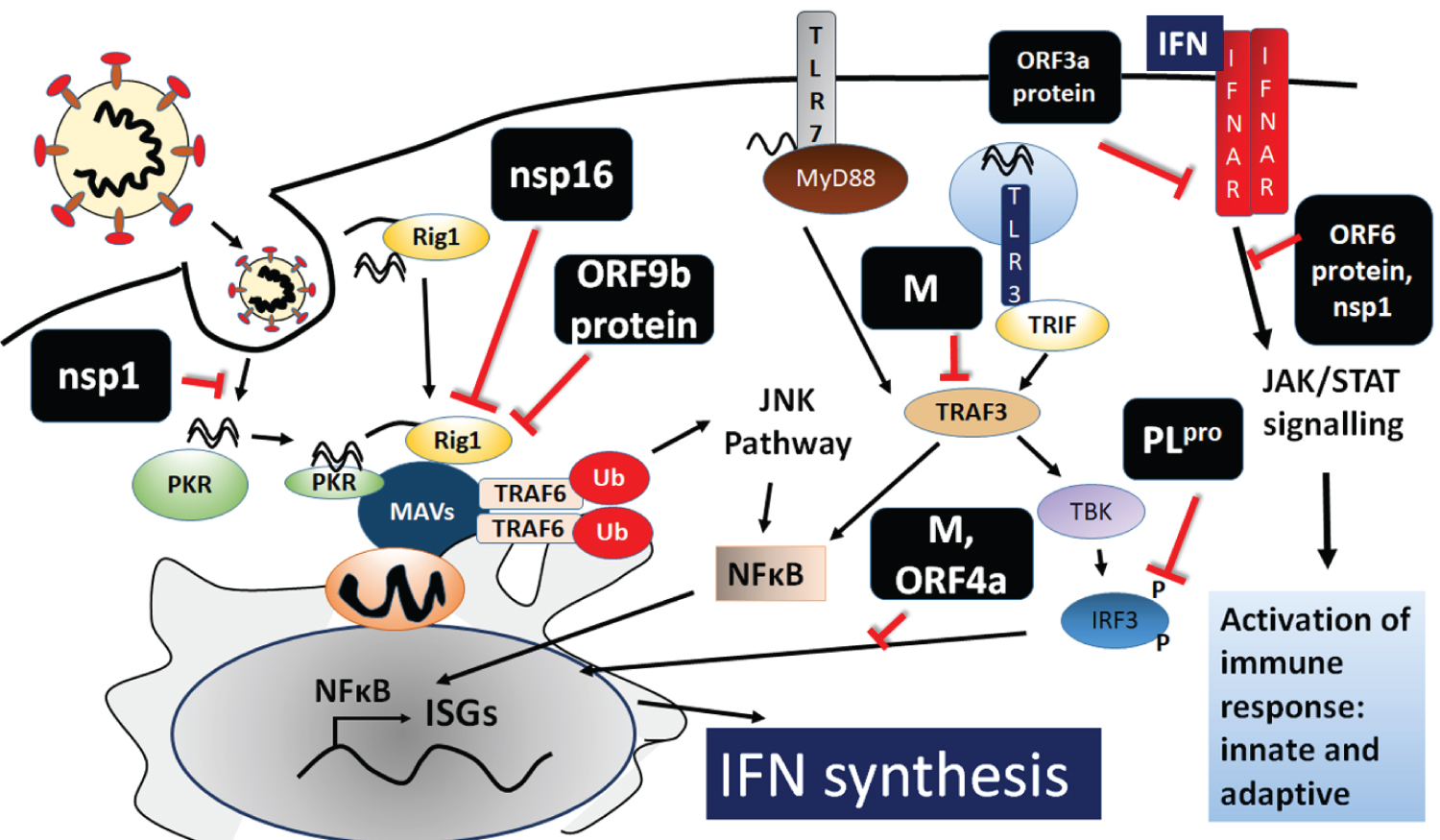
Figure 4: Innate Immune response in COVID-19.
Activation of interferon signalling system is depicted. The dsRNA (PAMP) intermediate during viral replication can trigger TLR3 (on endosomal membranes), RIG-1, MDA-5 and PKR all in the cytoplasm. GU-rich ssRNA can trigger TLR-7 which is present on endosomes and plasma membranes. TLR3 engages TRIF and TRAF3 which activate TBK, which phosphorylates IRF3 leading to its activation and translocation to the nucleus to activate transcription of interferons. Activation of TLR7 by MyD88 and TRAF3 activates IKKα resulting in the phosphorylation and activation of IRF-7, which too acts as a transcription factor similar to IRF3. Interferon genes are also activated by the NFκB pathway. MyD88 along with RIG-1 and MDA-5 converge on the MAVS (an outer mitochondrial membrane protein) coordinates activation of numerous TRAFs ultimately resulting in activation and translocation of NFκB to the nucleus. PKR also activates the NFκB pathway, apart from shutting down host mRNA translation. Upon release IFN signalling acts via JAK-STAT pathway. Various areas of interference by viral proteins (as predicted by studies in SARS-CoV and MERS) are also represented.
Evasion strategies by coronaviruses
Coronaviruses have also developed many ingenious means to tamper with the innate immune signalling response [68]. Primary evasion strategy by most RNA viruses replicating in the cytoplasm is to create a viroplasm of cellular membranes and hence sequester their genomic RNA from the cellular surveillance PRRs [69,70]. Furthermore, the N protein in SARS-CoV has been reported to sequester IFN-inducing RNA, while nsp1 mediates host mRNA degradation and blocks host mRNA translation and phosphorylation of STAT1 [71-73]. The M protein inhibits TRAF3/TBK1 complex formation whereas ORF9b product is known to induce proteosomal degradation of MAVs, TRAF3 and TRAF6 [74] (Figure 4). PLpro can inactivate IRF3, RIG-1, TBK1 and IRF3 [75] (Figure 4). Similar reports are also available about MERS in inhibiting various aspects of interferon synthesis and signalling molecules [76]. Furthermore, several protein products such as SARS-CoV ORF3a and ORF6 disrupt interferon-induced signalling by inducing degradation of the interferon α receptor (IFNAR) and inhibiting JAK/STAT signalling respectively [71]. JAK/STAT signalling is also disrupted by nsp1 of SARS-CoV [72]. Some zoonotic coronaviruses encode phosphodiesterase (PDE), which cleaves 2', 5'-oligoadenylate to block activation of cellular endo-ribonuclease RNase L, thereby inhibiting interferon-induced antiviral response, which is another intriguing feature that enhances virulence.
How does a cytokine storm happen?
The pathology of a cytokine storm [77] is a complicated event but we attempt to simplify it. As a result of interference at the first level of interferon signalling by the viral proteins, early stages of infection results in delayed release of cytokines and chemokines in the respiratory epithelial and dendritic cells [78,79]. During the later stages, low levels of interferons with higher levels of pro-inflammatory cytokines such as IL-β, TNF-α, and chemokines CCL-2, CCL-3 and CCL-5 [80-83] have been reported in SARS and MERS infections. This results in an accumulation of neutrophils and monocytes in the lung tissue and peripheral blood of the patient. An excess infiltration of these cells leads to lung injury, caused due to apoptosis of the infected cell. Furthermore, accumulated macrophages receive activating signals and produce more chemoattractants such as CCL-2, CCL-7, and CCL-12 that leads to a further accumulation of more immune competent cells. This results in an elevation of secreted TNF-α, IL-6, IL-1β [68,84] that induces apoptosis of T cells and damage to the pulmonary microvasculature, vascular leakage and alveolar edema.
A high prevalence of COVID-19 disease has been observed in immunocompromised population and patients with co-morbidities [85]. In case of obesity which is also a comorbid factor, it is proposed that cytokines are released upon inflammation leading to a severe condition in COVID-19 [86]. However, the underlying relation between comorbid factors and imbalanced immune response such as cytokine storm needs to be delved into further.
Adaptive Immune response to COVID-19: The adaptive or humoral response to SARS-CoV-2 forms the basis for the development of plasma therapy where plasma from cured patients are being administered to critical patients [87], as well as rapid ELISA based tests that don't require access to a sophisticated laboratory. The adaptive response develops as a result of signalling from the innate immune response arm but typically becomes prominent only after a few days of the onset of symptoms. In SARS-CoV, MERS as well as SARS-CoV-2, delayed and weak onset of the adaptive immune response has resulted in poor outcome [88,89]. Profiles of appearance of specific immunoglobulins from patients have revealed the appearance of acute phase antibodies, IgA, and IgM early after the onset of symptoms with later appearance of IgG. Consequently several ELISA detection kits are being developed, based on the detection of IgG [90].
There is a great volume of literature investigating whether a previous exposure to any of the human coronaviruses can generate immunity against SARS-CoV-2. Antibody profiling to check for cross reactivity between those generated against viral proteins of other coronaviruses with SARS-CoV-2 has almost consistently revealed mixed cross-reactivity results. While minimal cross reactivity has been observed between the S1 region of SARS and SARS-CoV-2 [91], a more conserved region of S2 has the possibility of more cross reactivity. Few other studies have also reported the effectiveness of neutralizing antibodies against SARS-CoV in inhibiting entry of SARS-CoV-2 [23]. Furthermore, antibodies against N protein has also shown certain amount of reactivity [90]. There have been reports on T-cell and B-cell response inducing epitope profiling and few epitopes are found to be common in the S and N proteins across SARS and SARS-CoV-2 [92]. Recently, a study suggests that an antibody cocktail consisting of four antibodies raised against spike variants have more effective neutralizing activity compared to individual antibody raised against specific spike protein of SARS-CoV-2. It prevents rapid mutational escape of the virus as seen with individual antibody [93].
Vaccine development and its effectiveness depend on the onset of adaptive immune response and hence ideally a perfect vaccine would induce long-lasting immunity against all coronaviruses. Hence, research has focused on a variety of targets, most common being the Spike protein as it is abundant and presented on the viral surface to be recognized early in infection. Furthermore, an effective vaccine against S protein could kick start the immune response instead of wait for the innate signalling arm to activate it. However, this depends on the level of conservation being maintained by Spike protein in all serotypes that are being generated and the epitopes being used in vaccine design.
An overview of antibody dependent enhancement (ADE) and its role in Coronavirus entry
Besides cross-species barriers, there are many other factors that throw challenges in developing effective vaccines and drug strategies against a zoonotic pathogen and coronavirus is no exception. Among other factors, antibody dependent enhancement of coronavirus entry is another issue which is of utmost importance in the fight against SARS-CoV-2. Over the last few years, scientists studied the mechanism of antibody dependent enhancement of pathogenicity and severity of Dengue infection (or any other Flaviviruses). Primary infection of dengue produces neutralizing antibodies against the same serotype. However, when the same patient if infected a second time by a different serotype, these neutralizing antibodies fail to protect but rather enhance the infection by many fold [94]. Such mechanism has also been observed in viruses like HIV and Ebola subverting antiviral immune response.
Antibody dependent enhancement or ADE is a phenomenon by which virus-specific antibodies accelerate the entry and/or replication of viruses into monocytes and granulocytes via interaction with FcR of IgG or complement receptors. It is facilitated either by uptake of antibody-virus complexes or by increasing viral nucleic acid and protein synthesis. This mechanism might also interfere with the cellular signalling pathways [94] or enhance stronger anti-inflammatory responses disproportionally [95]. Similarly, antibody dependent enhancement of infection has been observed in coronaviruses. Anti-sera against SARS-CoV spike enhances virus entry into FcR expressing cell [96]. This study reported that human pro-monocyte cell line (HL-CZ) which also expressed ACE2 was successfully infected by SARS-CoV in presence of monoclonal antibody targeted against the virus spike protein. Furthermore diluted anti-sera enhanced infection and induced apoptosis. It is important to note that virus-induced cytopathic effects and increased level of pro-inflammatory cytokines was also detected by quantitative RT-PCR. Additional studies indicate that immunization of cats with FCV spike worsens future infection due to ADE [97]. Another study experimentally demonstrated that neutralizing monoclonal antibody targeted against the RBD in spike protein of MERS and SARS coronaviruses catalyzes pseudovirus entry into FcR expressing host cells [98]. During ADE, the antibody likely binds RBD and leads to conformational changes, exposing S2' to protease cleavage. However, in case of SARS and MERS coronaviruses, receptors for ADE remain the same as for the normal viral entry, following ACE2 and DPP4-dependent viral entry. This indicates that neutralizing monoclonal antibody mediates ADE of coronavirus entry by acting as functional viral receptor mimic.
In line with humoral immune response, ADE becomes a major area of concern in vaccine design and antiviral drug therapy. At present there is no literature available on ADE of SARS-CoV-2. It can be perceived that SARS-CoV-2 might exhibit antibody dependent enhancement of viral entry and pathogenesis similar to its predecessors. Therefore, in this review we suggest to have research done in this area. If ADE is found in SARS-CoV-2, then strategy for antibody-dependent drug delivery needs to be modified accordingly. The current scenario where widespread vaccination is underway will also tell us about ADE in SARS-CoV-2 infections.
Current Therapeutic Approaches and Challenges
Repurposing drugs
With each passing day, the pandemic's toll on life and economy worldwide increases. Scientists across the globe are working tirelessly to find a cure, a solution to this pandemic. During this time of emergency, it is difficult to develop novel antiviral compound as drug testing takes time, thus prolonging start of a treatment regime. Therefore, researchers and public health agencies are screening the efficacy of existing drug combinations and therapies, those that have already been proved to be safe for other diseases. In this review a few most commonly used prophylactics and curatives have briefed in (Table 2) and (Table 3) respectively.
Table 2: Prophylactics. View Table 2
Table 3: Curatives. View Table 3
Race towards vaccine
The WHO has recognized about 48 candidates in the race towards development of an effective vaccine which are in the clinical evaluation and authorized globally. This article will aim to summarize some of the front runners in the following (Table 4).
Table 4: List of 10 frontline vaccine candidates. View Table 4
Since January 16, two COVID vaccine candidates Covishield and Covaxin were rolled out in India among the other frontline vaccines. Having vaccinated ~30 million people already, the country is now administering to people over 60 years of age [121] and those younger with comorbidities. Although both the vaccines have now accelerated the wheel, controversies over authorization of Covaxin, an inactivated whole virion vaccine continues. Also, the vaccines' effectiveness in high risk population have not been reported separately. Besides India, several other countries across the globe including Australia, South Africa, UK, Bangladesh, and Brazil authorized roll-out of Covishield (also known as ChAdOx1) since January 2021. However, several countries suspended the use of the vaccine after reports of blood clotting and thrombotic events began to circulate in March [122], which once again brings a pitfall in the fight against the virus and the pandemic. Other noteworthy vaccine candidates which are now in circulation are Comirnaty, mRNA-1273, SputnikV, COVID-19 vaccine Janssen and Coronavac. After its circulation, Comirnaty was claimed to be effective against the new variant B.1.1.7 but later it was proved wrong [123]. Even when we are chasing after the most promising treatment to fight against SARS-CoV-2, it is extremely important to keep in mind that unbiased use of so many therapeutics at the same time might complicate the treatment. When we are still at an early stage of vaccination, an upsurge of fresh infections in many countries including India, UK, Brazil, USA makes the fight against the pandemic even more cumbersome [124]. It might be possible that the pathogen has already developed a resistance against these therapeutics, gaining an advantage of high transmissibility and rapid disease progression. Therefore, more and more accurate clinical trials are needed before declaring the drug or vaccine, a promising one. Hence our fight against the disease seems to be a very tough one because as soon as we gain a miniscule edge against it, the virus seems to pose another challenge.
Plasma therapy
Since the MERS outbreak, plasma therapy has been the leading proposed treatment. Convalescent plasma (CP) therapy is a classical adaptive immunotherapy used to treat and prevent extremely infectious diseases like SARS, MERS, H1N1 swine flu [9]. In absence of a promising and approved specific antiviral drug against SARS-CoV-2, focus had shifted to alternate strategy to fight COVID-19, especially in severe patients. A study performed on a small group of patients demonstrated hopeful results. CP from recovered patients who had developed humoral immunity against SARS-CoV-2, with a large quantity of neutralizing antibodies raised against SARS-CoV-2 was administered to patients with severe disease. This therapy exhibited a promising change in the treatment procedure with little or no viral load in serum after CP transfusion, an increase in oxygen saturation and improvement of liver function and CRP level. The two key factors in plasma therapy were highlighted to be neutralizing antibody titre and treatment time point [88]. Another trial was conducted in China on a group of 5 critically ill COVID-19 patients with acute respiratory distress syndrome (ARDS). Although, viral load decreased after the standard antiviral treatment, it became undetectable only after plasma transfusion into recipients. Consistent with previous reports, there was a marked increase in neutralizing antibody titre, which highlights the possible clearance of the virus and clinical improvement [125]. Other countries like India and the UK is also continuing clinical trials with convalescent plasma therapy [126]. Central Drugs Standard Control Organization (CDSCO) of India also permitted Indian Council of Medical Research (ICMR) to conduct a series of trials of plasma therapy in certain states to fight against SARS-CoV-2 [127]. However, there are limitations to this therapy as well, which includes the small number of patients being treated and possible synergistic effect of other drugs with CP.
Discussion
Even after a year the entire world including more than 200 countries is still witnessing the rage of the pandemic COVID-19, a disease that has brought our life and livelihood to a halt or "new normal life" scientists are rigorously working to develop an effective antiviral therapeutic to defeat it. The fight has been multifaceted since the very beginning, whether fighting against economic recession or fight to adjust to the "new normal" life or fight to defeat the pandemic by designing effective therapeutics and vaccines. Some repurposed drugs have been approved but each drug has its own limitation. Therefore, science cannot rush; it can only make rational use of the potential drugs or vaccines. The world is in dire need of an effective vaccine. Although, the wait seems to be over since already a few vaccines have been authorized for use in several countries. However, it should be kept in mind that science meets with innumerable barriers, primarily due to lack of our understanding about the pathogen in depth, the complexities in human system, probable ADE of SARS-CoV-2 entry, cluster of mutated strains of the virus across the globe and lack of knowledge on virus life cycle, pathogenesis, high transmission rate and above all, time. Furthermore, unavailability of peer-reviewed data and clinical evidences on the already authorized vaccine candidates makes us repeatedly question on their efficiency and safety profile. Our only ray of hope lies in the fact this is not the first instance of a pathogen affecting humans and we have survived as a race either by developing therapeutics or by evolving our immunity to co-exist. It is not seldom that respiratory viruses have encountered humanity, and we shall overcome COVID-19 too taking all the lessons this pandemic has taught us and will be prepared for such pandemic outbreaks in coming days.
References
- Tyrrell DAJ, Bynoen ML (1965) Cultivation of a novel type of common-cold virus in organ cultures. Br Med J 1: 1467-1470.
- Drosten C, Günther S, Preiser W, et al. (2003) Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N Engl J Med 348: 1967-1976.
- Aly M, Elrobh M, Alzayer M, et al. (2017) Occurrence of the middle east respiratory syndrome coronavirus (MERS-cov) across the gulf corporation council countries: Four years update. PLoS One 12: 1-11.
- Zhou P, Yang XL, Wang XG, et al. (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579: 270-273.
- Su S, Wong G, Shi W, et al. (2016) Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol 24: 490-502.
- Jiang F, Deng L, Zhang L, et al. (2020) Review of the clinical characteristics of coronavirus disease 2019 (COVID-19). J Gen Intern Med 35: 1545-1549.
- Liu C, Zhou Q, Li Y, et al. (2020) Research and development on therapeutic agents and vaccines for COVID-19 and related human coronavirus diseases. ACS Cent Sci 6: 315-331.
- Raj VS, Mou H, Smits SL, et al. (2013) Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature 495: 251-254.
- De Wit E, Van Doremalen N, Falzarano D, et al. (2016) SARS and MERS: Recent insights into emerging coronaviruses. Nature Reviews Microbiology 14: 523-534.
- Wrapp D, Wang N, Corbett KS, et al. (2020) Cryo-EM structure of the 2019-ncov spike in the prefusion conformation. Science 367: 1260-1263.
- Báez-Santos YM, St John SE, Mesecar AD (2015) The SARS-coronavirus papain-like protease: Structure, function and inhibition by designed antiviral compounds. Antiviral Research 115: 21-38.
- Brian DA, Baric RS (2005) Coronavirus genome structure and replication. Curr Top Microbiol Immunol 287: 1-30.
- Perlman S, Netland J (2009) Coronaviruses post-SARS: Update on replication and pathogenesis. Nat Rev Microbiol 7: 439-450.
- Xia S, Liu M, C Wang, et al. (2020) Inhibition of SARS-cov-2 (previously 2019-ncov) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Res 2: 343-355.
- SKP L (2015) Structural insights into coronavirus entry-Elsevier Enhanced Reader.
- Eckerle LD, Becker MM, Halpin RA, et al. (2010) Infidelity of SARS-CoV Nsp14-exonuclease mutant virus replication is revealed by complete genome sequencing. PLoS Pathog 6: 1-15.
- Sevajol M, Subissi L, Decroly E, et al. (2014) Insights into RNA synthesis, capping, and proofreading mechanisms of SARS-coronavirus. Virus Res 194: 90-99.
- Schoeman D, Fielding BC (2019) Coronavirus envelope protein: Current knowledge. Virology Journal 16: 69.
- Torres J, Maheswari U, Parthasarathy K, et al. (2007) Conductance and amantadine binding of a pore formed by a lysine-flanked transmembrane domain of SARS coronavirus envelope protein. Protein Sci 16: 2065-2071.
- Torres J, Surya W, Li Y, et al. (2015) Protein-protein interactions of viroporins in coronaviruses and paramyxoviruses: New targets for antivirals?. Viruses 7: 2858-2883.
- Wan Y, Shang J, Graham R, et al. (2020) Receptor recognition by novel coronavirus from Wuhan: An analysis based on decade-long structural studies of SARS. J Virol 94.
- Donoghue M, Hsieh F, Baronas E, et al. (2000) Ultrarapid communication A novel angiotensin-converting enzyme – related to angiotensin 1-9. Circ Res 87: e1-e9.
- Hoffmann M, Kleine-Weber H, Schroeder S, et al. (2020) SARS-cov-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Elsevier BV 181: 271-280.
- Xia S, Yan L, Xu W, et al. (2019) A pan-coronavirus fusion inhibitor targeting the HR1 domain of human coronavirus spike.
- Yan R, Zhang Y, Li Y, et al. (2020) Structural basis for the recognition of the SARS-cov-2 by full-length human ACE2. Science 367: 1444-1448.
- Simmons G, Gosalia DN, Rennekamp AJ, et al. (2005) Inhibitors of cathepsin L prevent severe acute respiratory syndrome coronavirus entry. Proc Natl Acad Sci 102: 11876-11881.
- Ou X, Liu Y, Lei X, et al. (2020) Characterization of spike glycoprotein of SARS-cov-2 on virus entry and its immune cross-reactivity with SARS-cov. Nat Commun 11: 1620.
- Wang Q, Zhang Y, Wu L, et al. (2020) Structural and functional basis of SARS-cov-2 entry by using human ACE2. Cell 181: 894-904.
- Hoffmann M, Kleine-Weber H, Pöhlmann S (2020) A Multibasic cleavage site in the spike protein of SARS-cov-2 is essential for infection of human lung cells. Mol Cell 78: 779-784.
- Doremalen van N, Bushmaker T, Morris DH, et al. (2020) Aerosol and surface stability of SARS-CoV-2 as compared with SARS-cov-1. N Engl J Med 382: 1564-1567.
- Zhang T, Wu Q, Hang Z (2020) Probable pangolin origin of SARS-cov-2 associated with the COVID-19 outbreak. Curr Biol 30: 1346-1351.
- Millet JK (2017) Physiological and molecular triggers for SARS-CoV membrane fusion and entry into host cells _ Elsevier Enhanced Reader.
- Stadler K, Masignani V, Eickmann M, et al. (2003) SARS - beginning to understand a new virus. Nat Rev Microbiol 1: 209-218.
- Sing F (2014) Coronavirus-induced ER stress response and its involvement in regulation of coronavirus–host interactions _ elsevier enhanced reader.
- Zheng J (2020) SARS-cov-2: an Emerging coronavirus that causes a global threat. Int J Biol Sci 16: 1678-1685.
- Zhang J, Jia W, Zhu J, et al. (2020) Insights into the cross-species evolution of 2019 novel coronavirus. J Infect 80: 671-693.
- Yuen KS, Ye ZW, Fung SY, et al. (2020) SARS-CoV-2 and COVID-19: The most important research questions. Cell Biosci 10: 1-5.
- Zhai X, Sun J, Yan Z, et al. (2020) Comparison of SARS-cov-2 spike protein binding to ACE2 receptors from human, pets, farm animals, and putative intermediate hosts. J Virol.
- Gretebeck LM, Subbarao K, (2015) Animal models for SARS and MERS coronaviruses. Curr Opin Virol 13: 123-129.
- Kim YI, Kim SG, Kim SM, et al. (2020) Infection and rapid transmission of SARS-cov-2 in ferrets. Cell Host Microbe 1-6.
- Graham RL, Baric RS (2010) Recombination, Reservoirs, and the modular spike: mechanisms of cronavirus cross-species transmission. J Virol 84: 3134-3146.
- Woo PCY, SKP Lau, Huang Y (2009) Coronavirus diversity, phylogeny and interspecies jumping. Exp Biol Med 234: 1117-1127.
- Stanhope MJ, Brown JR, Amrine-Madsen H (2004) Evidence from the evolutionary analysis of nucleotide sequences for a recombinant history of SARS-CoV. Infect Genet Evol 4: 15-19.
- Khailany RA, Safdar M, Ozaslan M (2020) Genomic characterization of a novel SARS-CoV-2. Gene Reports 19: 100682.
- F Diego (2017) Molecular Evolution of Human Coronavirus Genomes _ Elsevier Enhanced Reader.
- Qu XX, Hao P, Song XJ, et al. (2005) Identification of two critical amino acid residues of the severe acute respiratory syndrome coronavirus spike protein for its variation in zoonotic tropism transition via a double substitution strategy. J Biol Chem 280: 29588-29595.
- Forster P, Forster L, Renfrew C (2020) Phylogenetic network analysis of SARS-CoV-2 genomes. Proc Natl Acad Sci 117: 9241-9243.
- Zhang, (2020) Genomic variations of SARS-CoV-2 suggest multiple outbreak sources of transmission.
- C Yin (2020) Genotyping coronavirus SARS-CoV-2: Methods and implications. 19: 1-12.
- Ortega JT, Serrano ML, Pujol FH, et al. (2020) Original article : Role of changes in sars-cov-2 spike protein in the interaction with the human ace2 receptor . EXCLI J 19: 410-417.
- Stefanelli P, Faggioni G, Presti A Lo, et al. (2020) Whole genome and phylogenetic analysis of two SARS-CoV-2 strains isolated in Italy in January and February 2020: additional clues on multiple introductions and further circulation in Europe. Eurosurveillance 25: 1-5.
- T Phan (2020) Genetic diversity and evolution of SARS-CoV-2. Infect Genet Evol 81: 104260.
- Benvenuto D, Angeletti S, Giovanetti M, et al. (2020) Evolutionary analysis of SARS-2-cov: How mutation of non-structural protein 6 (NSP6) could affect viral autophagy. SSRN ElectronJ 10: 3-6.
- Joshi, Paul S (2020) Phylogenetic analysis of the novel coronavirus reveals important variants in indian strains.
- Leung K, Pei Y, Leung GM, et al. (2020) Empirical transmission advantage of the D614G mutant strain of SARS-CoV-2 (preprint).
- Hou YJ, Chiba S, Halfmann P, et al. (2020) SARS-CoV-2 D614G variant exhibits efficient replication ex vivo and transmission in vivo. 8499: 1-11.
- Plante JA, Liu Y, Liu J, et al. (2020) Spike mutation D614G alters SARS-CoV-2 fitness. Nature 592: 116-121.
- Volz E, Hill V, McCrone JT, et al. (2020) Evaluating the effects of SARS-CoV-2 Spike mutation D614G on transmissibility and pathogenicity. Cell 184: 64-75.e11.
- (2021) New COVID-19 Variants | CDC.
- (2021) Threat Assessment Brief: Rapid increase of a SARS-CoV-2 variant with multiple spike protein mutations observed in the United Kingdom.
- Channappanavar R, Perlman S (2017) Pathogenic human coronavirus infections: causes and consequences of cytokine storm and immunopathology. Springer 39: 529-539.
- McGonagle D, Sharif K, O’Regan A, et al. (2020) The Role of Cytokines including Interleukin-6 in COVID-19 induced Pneumonia and Macrophage Activation Syndrome-Like Disease. Autoimmunity Reviews Elsevier 19: 102537.
- Ye Q, Wang B, Mao J (2020) The pathogenesis and treatment of the `Cytokine Storm’ in COVID-19. J Infect 80: 607-613.
- Felsenstein S, Herbert JA, McNamara PS, et al. (2020) COVID-19: Immunology and treatment options. Clin Immunol 215: 108448.
- Li G, Fan Y, Lai Y, et al. (2020) Coronavirus infections and immune responses. Journal of Medical Virology 92: 424-432.
- Del Prete, Baerwald C, Russo RC, et al. (2019) Cytokine Storm in COVID-19: ‘When you come out of the storm, you won’t be the same person who walked in. Front Immunol 11: 2132.
- Billiau A (2021) Interferon beta in the cytokine network: An anti-inflammatory pathway. Mult Scler 1: S2-S4.
- Totura L, Baric RS (2012) SARS coronavirus pathogenesis: Host innate immune responses and viral antagonism of interferon. Current Opinion in Virology 2: 264-275.
- Versteeg GA, Bredenbeek PJ, SHE van den Worm, et al. (2007) Group 2 coronaviruses prevent immediate early interferon induction by protection of viral RNA from host cell recognition. Virology 361: 18-26.
- van Hemert MJ, van den Worm SHE, Knoops K, et al. (2008) SARS-Coronavirus Replication/Transcription Complexes Are Membrane-Protected and Need a Host Factor for Activity In Vitro. PLoS Pathog 4: 1000054.
- Kopecky-Bromberg SA, Martínez-Sobrido L, Frieman M, et al. (2007) Severe Acute Respiratory Syndrome Coronavirus Open Reading Frame (ORF) 3b, ORF 6, and Nucleocapsid Proteins Function as Interferon Antagonists. J Virol 81: 548-557.
- Wathelet MG, Orr M, Frieman MB, et al. (2007) Severe Acute Respiratory Syndrome Coronavirus Evades Antiviral Signaling: Role of nsp1 and Rational Design of an Attenuated Strain. J Virol 81: 11620-11633.
- Huang C, Lokugamage KG, Rozovics JM, et al. (2011) SARS coronavirus nsp1 protein induces template-dependent endonucleolytic cleavage of mRNAs: Viral mRNAs are resistant to nsp1-induced RNA cleavage. PLoS Pathog 7: 1002433.
- Siu KL, Kok KH, Poon VKM, et al. (2009) Severe acute respiratory syndrome coronavirus M protein inhibits type I interferon production by impeding theformation of TRAF3•TANK•TBK1/IKKε complex. J Biol Chem 284: 16202-16209.
- Mielech M, Kilianski A, Baez-Santos YM, et al. (2014) MERS-CoV papain-like protease has deISGylating and deubiquitinating activities. Virology 450-451: 64-70.
- Yang Y, Zhang L, Genget H, et al. (2013) The structural and accessory proteins M, ORF 4a, ORF 4b, and ORF 5 of Middle East respiratory syndrome coronavirus (MERS-CoV) are potent interferon antagonists. Protein Cell 4: 951-961.
- Tisoncik JR, Korth MJ, Simmons CP, et al. (2012) Into the Eye of the Cytokine Storm. Microbiol Mol Biol Rev 76: 16-32.
- Faure E, Poissy J, Goffard A, et al. (2014) Distinct Immune Response in Two MERS-CoV-Infected Patients: Can We Go from Bench to Bedside?. PLoS One 9: e88716.
- Kindler E, Thiel V (2016) SARS-CoV and IFN: Too Little, Too Late. Cell Host and Microbe 19: 139-141.
- Cheung CY, Poon LL, Ng IH, et al. (2005) Cytokine Responses in Severe Acute Respiratory Syndrome Coronavirus-Infected Macrophages In Vitro: Possible Relevance to Pathogenesis. J Virol 79: 7819-7826.
- Chien JY, Hsueh PR, Cheng WC, et al. (2006) Temporal changes in cytokine/chemokine profiles and pulmonary involvement in severe acute respiratory syndrome. Respirology 11: 715-722.
- Law HKW, Cheung CY, Ng HY, et al. (2005) Chemokine up-regulation in SARS-coronavirus-infected, monocyte-derived human dendritic cells. Blood 106: 2366-2374.
- Lau SKP, Candy CY Lau, Kwok-Hung Chan, et al. (2013) Delayed induction of proinflammatory cytokines and suppression of innate antiviral response by the novel Middle East respiratory syndrome coronavirus: Implications for pathogenesis and treatment. J Gen Virol 94: 2679-2690.
- Thiel V, Weber F (2008) Interferon and cytokine responses to SARS-coronavirus infection. Cytokine and Growth Factor Review 19: 121-132.
- Wang L, He W, Yu X, et al. (2020) Coronavirus disease 2019 in elderly patients: Characteristics and prognostic factors based on 4-week follow-up. J Infect 80: 639-645.
- Prompetchara E, Ketloy C, Palaga T (2020) Allergy and Immunology Immune responses in COVID-19 and potential vaccines: Lessons learned from SARS and MERS epidemic.
- Duan K, Liu B, Cesheng Li, et al. (2020) Effectiveness of convalescent plasma therapy in severe COVID-19 patients. Proc Natl Acad Sci 117: 9490-9496.
- Liu W, Fontanet A, Zhang P-H, et al. (2006) Two‐Year Prospective Study of the Humoral Immune Response of Patients with Severe Acute Respiratory Syndrome. J Infect Dis 193: 792-795.
- Liu WJ, Zhao M, Liu K, et al. (2017) T-cell immunity of SARS-CoV: Implications for vaccine development against MERS-CoV. Antiviral Research 137: 82-92.
- Guo L, Ren L, Yang S, et al. (2020) Profiling Early Humoral Response to Diagnose Novel Coronavirus Disease (COVID-19).
- Walls C, Park YJ, Tortorici MA, et al. (2020) Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein.Cell 181: 281-292.
- Ahmed SF, Quadeer AA, McKay MR (2020) Preliminary identification of potential vaccine targets for the COVID-19 Coronavirus (SARS-CoV-2) Based on SARS-CoV Immunological Studies. Viruses 12: 254.
- Baum A, Fulton BO, Wloga E, et al. (2020) Antibody cocktail to SARS-CoV-2 spike protein prevents rapid mutational escape seen with individual antibodies. Science 369: 1014-1018.
- Flipse J, Diosa-Toro MA, Hoornweg TE, et al. (2016) Antibody-Dependent Enhancement of Dengue Virus Infection in Primary Human Macrophages; Balancing Higher Fusion against Antiviral Responses. Sci Rep 6: 1-13.
- Thacker B (2003) PRRS virus infection and disease. PRRS Compend 16: 11-16.
- Wang SF (2014) ADE of SARS.
- Corapi WV, Olsen CW, Scott FW (1992) Monoclonal Antibody Analysis of Neutralization and Antibody-Dependent Enhancement of Feline Infectious Peritonitis Virus. J Virol 66: 6695-6705.
- Wan Y, Shang J, Sun S, et al. (2019) Molecular Mechanism for Antibody-Dependent Enhancement of Coronavirus Entry. J Virol 94: e02015-e02019.
- Vincent MJ, Bergeron E, Benjannet S, et al. (2005) Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virol J 2: 1-10.
- Liu J, Cao R, Xu M, et al. (2020) Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting SARS-CoV-2 infection in vitro. Cell Discov 6: 16.
- Gautret P, Lagier JC, B Doudier, et al. (2020) Hydroxychloroquine and azithromycin as a treatment of COVID-19: results of an open-label non-randomized clinical trial. Int J Antimicrob Agents 56: 105949.
- Valle C, Martin B, Touret F, et al. (2020) Drugs against SARS-CoV-2: What do we know about their mode of action? Rev Med Virol 30: 1-10.
- Wang X, Cao R, Zhang H, et al. (2020) The anti-influenza virus drug, arbidol is an efficient inhibitor of SARS-CoV-2 in vitro. Cell Discov 6: 4-8.
- Deng L, Li C, Zeng Q, et al. (2020) Arbidol combined with LPV/r versus LPV/r alone against Corona Virus Disease 2019: A retrospective cohort study. J Infect 81: e1-e5.
- Sheahan TP, Sims AC, Graham RL, et al. (2017) Broad-spectrum antiviral GS-5734 inhibits both epidemic and zoonotic coronaviruses. Sci Transl Med 9: eaal3653.
- Cao Yc, Deng QX, Dai SX, et al. (2020) Remdesivir for severe acute respiratory syndrome coronavirus 2 causing COVID-19: An evaluation of the evidence. Travel Medicine and Infectious Disease 35: 101647.
- Herper M (2020) Inside the NIH’s controversial decision to stop its big remdesivir study. STAT PARS International.
- Tu YF, Chien CS, Yarmishyn AA, et al. (2020) A review of sars-cov-2 and the ongoing clinical trials. Int J Mol Sci 21: 2657.
- Cheng CY (2020) Lopinavir-ritonavir did not shorten the duration of SARS CoV-2 shedding in patients with mild pneumonia in Taiwan -Elsevier Enhanced Reader.pdf.
- Matthay MA, Thompson BT (2020) Dexamethasone in hospitalised patients with COVID-19: Addressing uncertainties. Lancet Respir Med 19-20.
- (2020) Oxford COVID-19 vaccine begins human trial stage. Research University of Oxford.
- Stage C, et al. (2020) DRAFT landscape of COVID-19 candidate vaccines.
- Cadila Z, Biologicals N, Zycov-d W (2020) Zydus Cadila Announces Completion of Dosing in Phase I Clinical Trial of ZyCoV-D. 63.
- Mulligan MJ, Lyke KE, Kitchin N, et al. (2020) Phase I / II study of COVID-19 RNA vaccine BNT162b1 in adults. Nature 586: 589-593.
- Sahin U, Muik A, Türeci O, et al. (2020) COVID-19 vaccine BNT162b1 elicits human antibody and TH1T cell responses. Nature 586: 594-599.
- Makhene M, Anderson EJ, Rouphael NG, et al. (2020) Safety and Immunogenicity of SARS-CoV-2 mRNA-1273 Vaccine in older adults. N Engl J Med 1-12.
- Keech C, Albert G, Cho I, et al. (2020) Phase 1-2 Trial of a SARS-CoV-2 Recombinant Spike Protein Nanoparticle Vaccine. N Engl J Med 1-13.
- Babira VF, Borisevich SV, Naroditsky BS, et al. (2020) Safety and immunogenicity of an rAd26 and rAd5 vector-based heterologous prime-boost COVID-19 vaccine in two formulations : Two open , non-randomised phase 1 / 2 studies from Russia. Lancet 396: 887-897.
- Miller A, Reandelar MJ, Fasciglione K, et al. (2020) Correlation between universal BCG vaccination policy and reduced morbidity and mortality for COVID-19: An epidemiological study.
- Berg MK, Yu Q, Salvador CE, Melani I, Kitayama S (2020) Mandated Bacillus Calmette-Guérin (BCG) vaccination predicts flattened curves for the spread of COVID-19. Sci Adv 6: 1-8.
- (2021) Covishield and Covaxin: What we know about India’s Covid-19 vaccines.
- Paul-Ehrlich-Institut News (2021) The Paul-Ehrlich-Institut informs-Temporary Suspension of Vaccination with COVID-19 Vaccine AstraZeneca.
- Liu Y, Liu J, Xia H, et al. (2021) Neutralizing activity of BNT162b2-Elicited Serum - Preliminary Report. N Engl J Med.
- (2021) Covid-19 update: India records biggest spike in new cases in over 100 days.
- Shen C, Wang Z, Zhao F, et al. (2020) Treatment of 5 Critically Ill Patients With COVID-19 With Convalescent Plasma. JAMA 29: 1-8.
- (2020) UK approves clinical trial of plasma therapy for Covid-19.
- (2020) India’s CDSCO permits convalescent plasma trial for Covid-19.
Keywords
INDEXING
PARTNERS









Table 1: Comparative features between SARS-CoV and CoV-2. View Table 1